Automated Ingestion
Automated pipelines continuously ingest publications and abstracts from repositories like PubMed and major medical conferences.
Client: A life sciences intelligence platform scaling its
clinical data offerings.
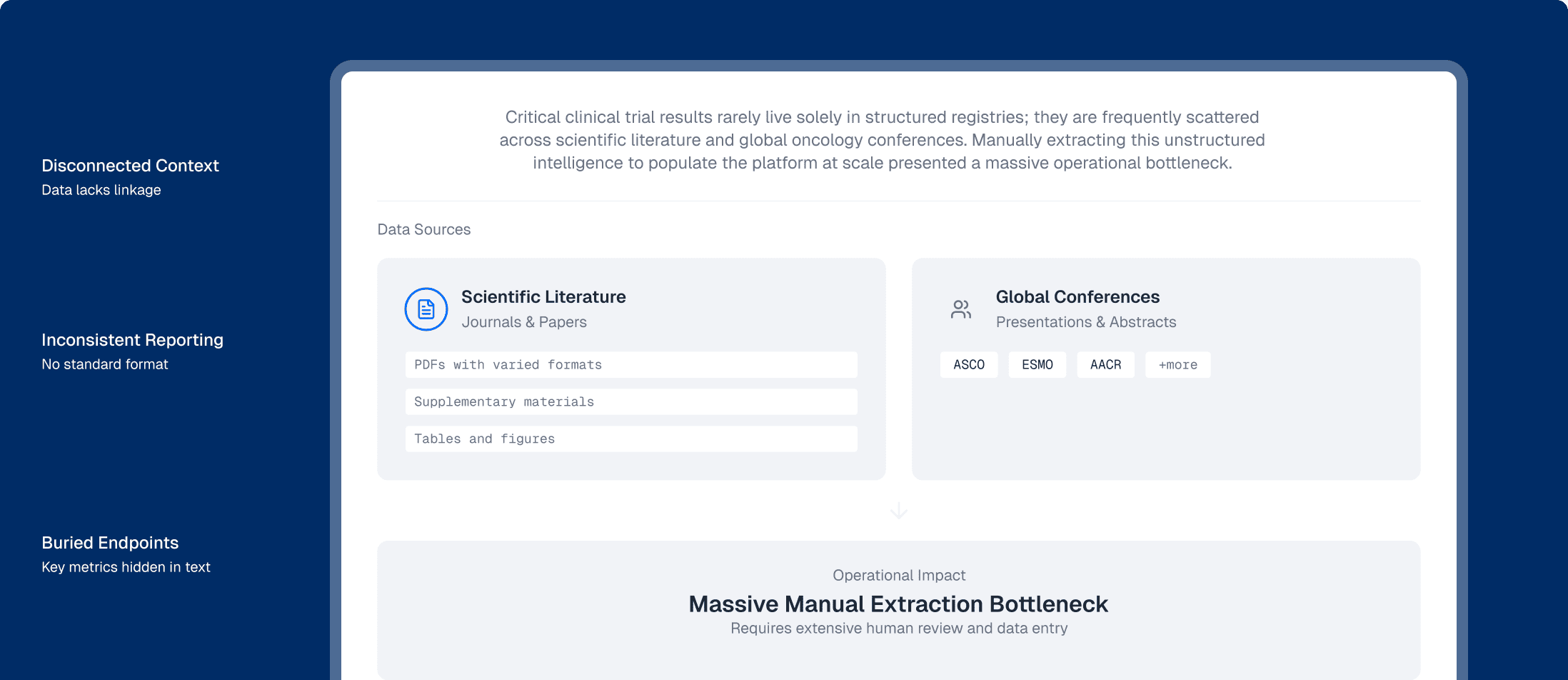
Across the life sciences industry, critical clinical trial results rarely live solely in structured registries; they are frequently scattered across scientific literature and global oncology conferences (such as ASCO, ESMO, and AACR). For the client, manually extracting this unstructured intelligence to populate their platform at scale presented a massive operational bottleneck:
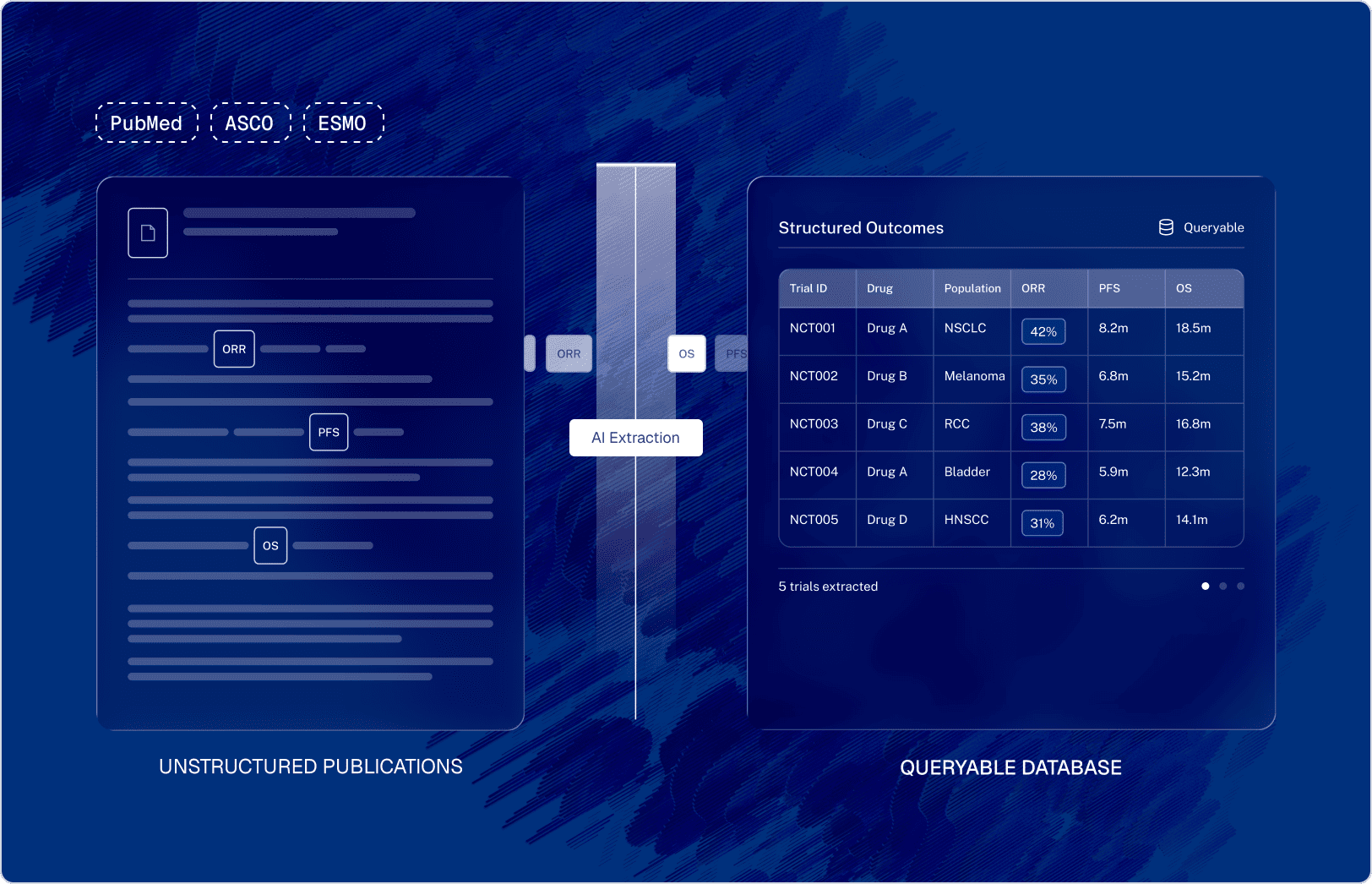
To industrialize the client's literature review process, Sogody deployed containerized AI pipelines designed specifically to ingest, extract, and standardize complex biomedical publications.
Automated pipelines continuously ingest publications and abstracts from repositories like PubMed and major medical conferences.
Embedded large language models process the unstructured text to automatically identify and extract trial references (such as NCT IDs), specific drug interventions, and detailed patient populations or sub-groups.
The AI agents don't just extract text; they normalize complex trial outcomes. Disparate mentions of survival rates and tumor responses are mapped to a standardized hierarchy of clinical endpoints (like ORR and PFS), ensuring consistent measurement definitions across all extracted data.
The extracted intelligence passes through a rigorous quality gate. Data points that fall outside expected confidence thresholds or present anomalies are flagged and diverted to a "Human Review" loop, ensuring that the system maintains strict scientific accuracy before data is released to the platform.
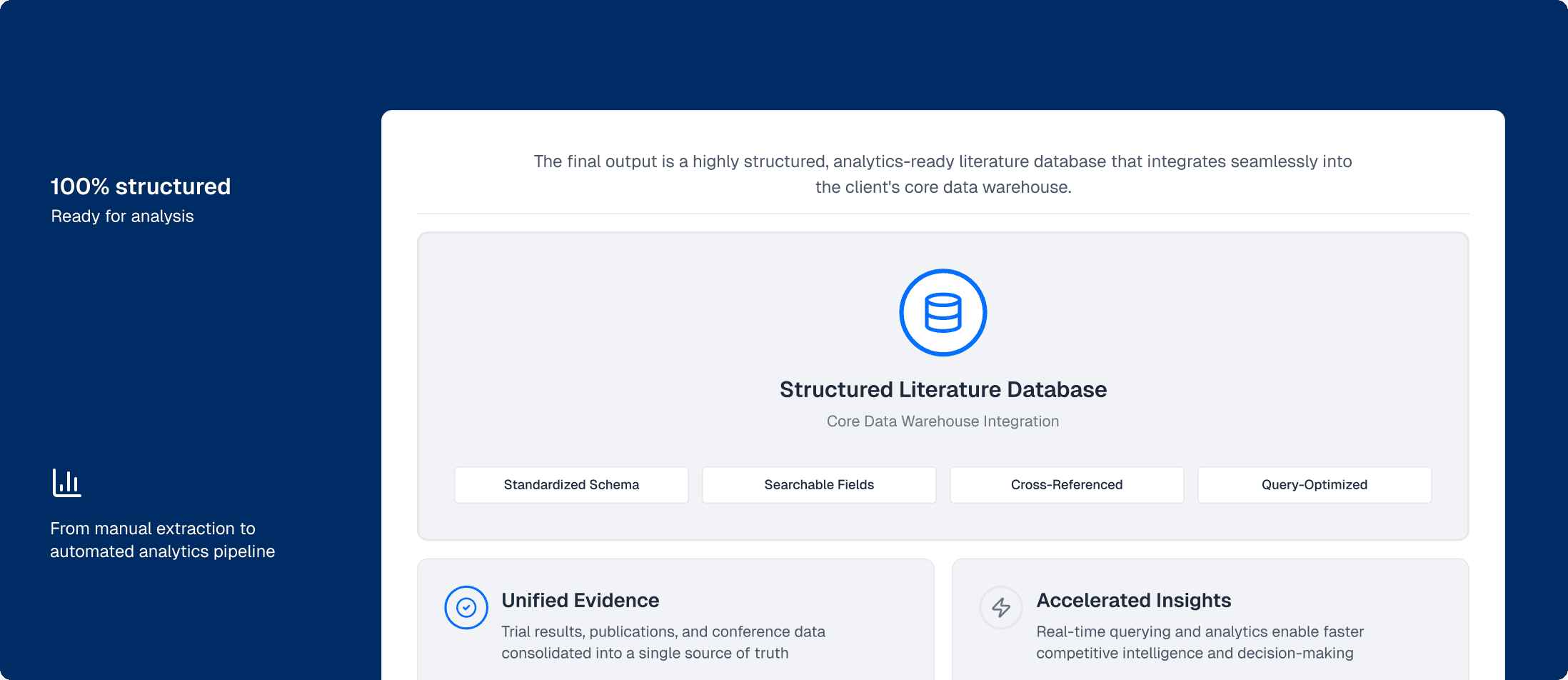
The final output is a highly structured, analytics-ready literature database that integrates seamlessly into the client's core data warehouse.